I've spend much of my time since Monday working to get the agent-based model I'm currently messing around with (version 3 of ForagerNet3_Demography) to work in version 2.4 of Repast Simphony. Something went awry with my previous installation (2.0) and it made sense to start trying to fix things with the most current version. After many frustrating loops of install/uninstall/reinstall, I finally got a good installation of 2.4 and got my source files moved in and working.
One thing that I wasn't able to make functional in the last version was the batch mode. Batch mode lets you run jobs in batches, speeding up computation by farming out the computational workload to different processors or machines, etc. (doing runs in parallel rather than serial). It wasn't too troublesome that I couldn't get the embedded batch mode working before, as the model I'm working with isn't terribly demanding. There are bigger things coming down the road, however, so I was happy to see the batch mode appeared to be active and ready to go with the new installation.
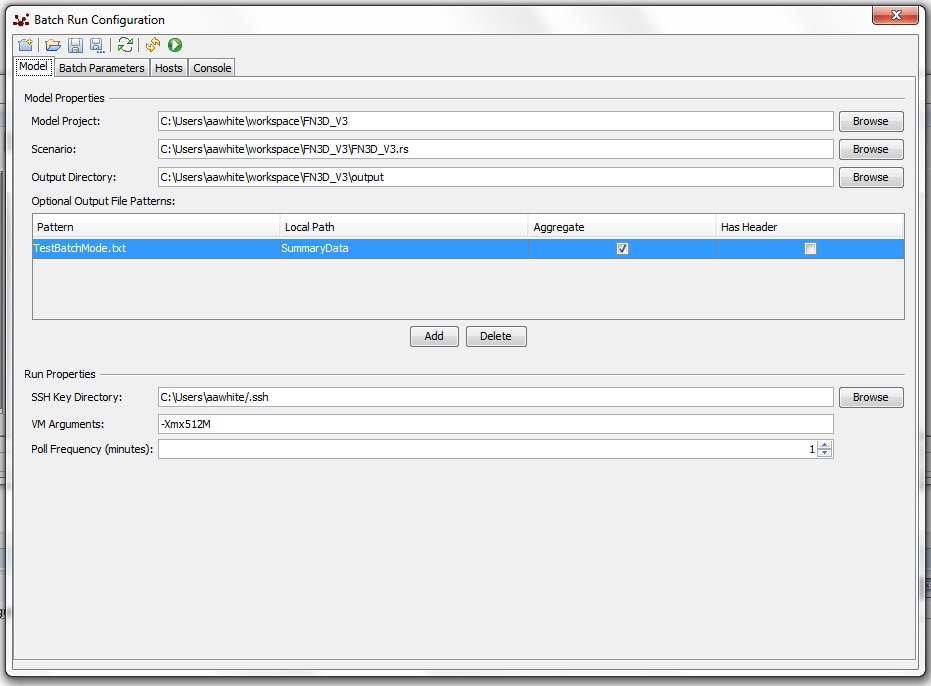
I spent most of yesterday trying to figure out how to use the batch mode while preserving the data output format that I've created on my own and that works for me. The way I've written the model, it produces a summary data file at the end of each run. The file (just a simple .txt file that I can open in a spreadsheet) preserves information on all the parameter settings used in the run as well as data on the outcomes. In the Repast/Eclipse batch mode, however, that .txt file wasn't getting produced. I was frustrated to learn that I apparently had to redo all the data capturing in the model so that the software could use something called a "file sink" to collect information. I was then relieved to learn that there was a workaround to using the "embedded system:" you can specify an "Optional Output File Pattern:"
Repast seems to be picky about the settings in the "batch_params.xml" and "parameters.xml" files, wanting them to match. I get null pointer exceptions when I clear everything but the random seed out of the files, so I just left a parameter in to keep the software happy (it does nothing, as it's a constant and doesn't affect any of the settings in the model).
I'm still testing out the batch mode and the model to make sure everything is working properly. My plan then is to modify version 3 to create version 4, which will include some different mortality settings (which, in turn, will necessitate rewriting some of the code that calculates age-specific mortality outcomes). After all that's done, I'm planning on doing some new modeling work to formally follow up on some preliminary observations I presented at the SAA meetings a few years ago.


 RSS Feed
RSS Feed
